Faithful and Stable Neuron Explanations for Trustworthy Mechanistic Interpretability
Abstract
- We observe that neuron identification (NI) can be viewed as the inverse process of machine learning, providing a unified theoretical lens for analyzing explanation quality.
- We derive generalization bounds for popular similarity metrics (accuracy, AUROC, IoU) to guarantee faithfulness of the identified concept to the neuron's true behavior.
- We propose a Bootstrap Explanation (BE) method that quantifies stability of explanations across probing datasets and produces concept prediction sets with guaranteed coverage probability.
Overview
(i) Introduction
Recent progress in mechanistic interpretability has made it possible to automatically name or describe internal neurons using curated datasets, multimodal embeddings, or language model summarization. Yet two questions remain central:
- Faithfulness: does the selected concept accurately reflect the neuron's behavior?
- Stability: would we obtain the same concept if we probed with a different dataset?
This paper answers both questions under a unified perspective: neuron identification can be cast as selecting \( \hat c \in \mathcal{C} \) that maximizes an empirical similarity \( \hat{\mathrm{sim}}(f, c; D_{\text{probe}}) \) between neuron output \( f(x) \) and concept activation \( c(x) \). Because this is structurally identical to empirical risk minimization, we can adapt classical generalization arguments to bound the deviation \( |\hat{\mathrm{sim}} - \mathrm{sim}| \) and thus obtain high-probability guarantees on faithfulness. For stability, we adopt a bootstrap ensemble approach that quantifies uncertainty in concept selection and produces prediction sets with statistical guarantees.
(ii) Formalizing Neuron Identification
A neuron (or feature direction, SAE feature, linear probe, TCAV direction) is modeled as a function \( f: \mathcal{X} \to \mathbb{R} \) mapping inputs (e.g. images) to activations. A concept is modeled as \( c: \mathcal{X} \to [0,1] \), the probability that the concept is present. Given a concept set \( \mathcal{C} \), a similarity functional \( \mathrm{sim}(f, c) \), and a probing dataset \( D_{\text{probe}} = \{x_i\}_{i=1}^n \), neuron identification is
$$ \hat c = \arg\max_{c \in \mathcal{C}} \ \hat{\mathrm{sim}}(f, c; D_{\text{probe}}), $$where \( \hat{\mathrm{sim}}\) denotes the empirical estimation of similarity on the probing dataset. Different similarity choices (accuracy, AUROC, IoU, recall, precision) can all be placed in this form, which gives a single theoretical lens for comparing them.
(iii) Connection to Machine Learning
Our key observation is that neuron identification objective is similar to that of supervised learning: $$ \hat{h} = \arg \min_{h \in H} \quad \hat{L}(h; D_{\text{train}}). $$ Comparing two objectives, we find they are highly similar, except that instead of finding a model to fit human labels, neuron identification finds a human concept to fit a model (the neuron). This analogy allows us to adapt classical learning-theoretic tools to analyze neuron identification.
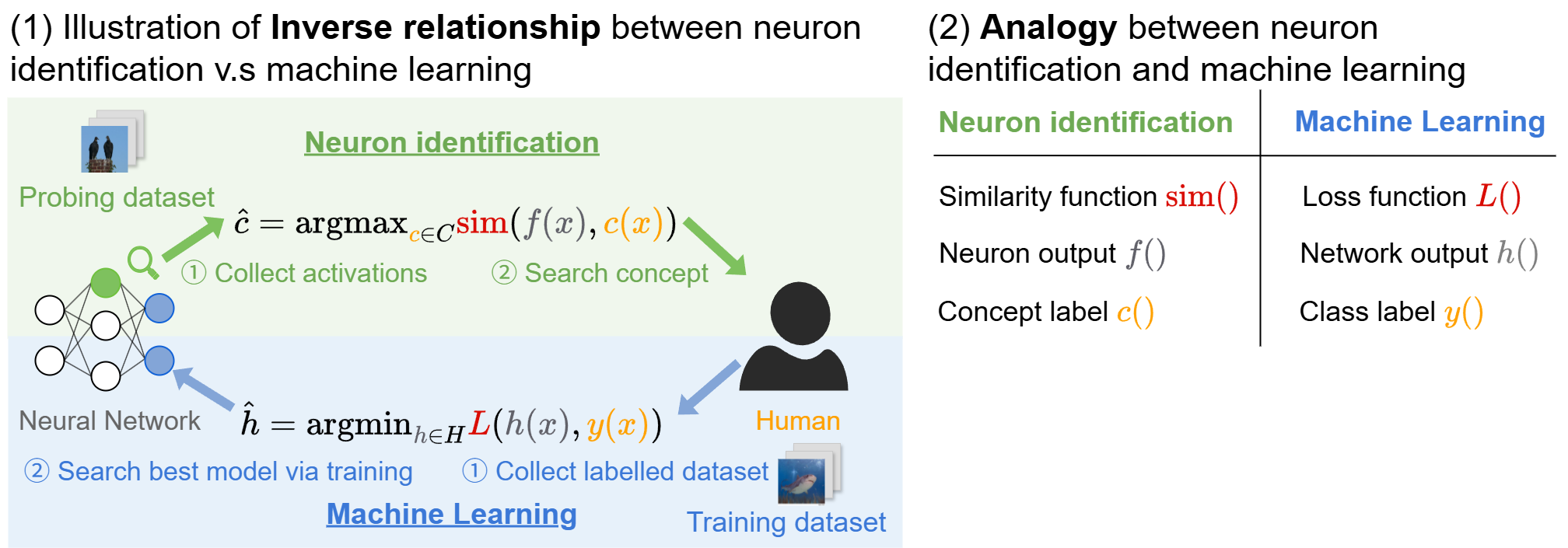
Figure 1: Neuron identification is viewed as the inverse of supervised learning: instead of finding a model to match human labels, we find a human concept to match a neuron. This perspective enables learning-theoretic guarantees.
Contribution #1: Theoretical Guarantees for Explanation Faithfulness
Define the generalization gap \[ g(D_{\text{probe}}, \mathcal{C}, f) \triangleq \sup_{c \in \mathcal{C}} \big(\hat{\mathrm{sim}}(f, c; D_{\text{probe}}) - \mathrm{sim}(f, c)\big). \] Under mild assumptions that
- The concept set \( \mathcal{C} \) is finite;
- The probing dataset \(D_{\text{probe}}\) is sampled i.i.d. from the true distribution;
- The similarity metric is bounded within [0, 1];
As a corollary, the empirically best concept \( \hat c \) is within \( 2r(\cdot) \) of the true best concept \( c^\star \), showing that the reported explanation is approximately optimal on the true distribution provided the probing dataset is large enough and the concept set is not too large. This is the first such guarantee tailored to neuron identification.
(i) Convergence for popular metrics
To understand and compare popular similarity metrics, we instantiate the general result for several widely used similarity metrics by calculating their convergence rates \( r(\cdot) \). In general, most metrics have a convergence rate of \( O(1/\sqrt{n}) \) where \( n \) is the probing dataset size, but differ in constants:
- Accuracy: Hoeffding-style bound, converges fast in practice.
- AUROC: depends on concept frequency \( \rho(c) \); rare concepts converge slowly.
- IoU, precision, recall: can be seen as accuracy on a restricted subset, so the effective sample size becomes smaller and convergence is slower.
(ii) Simulation study
To verify our theory, we simulate neuron outputs and concepts with controlled similarity levels, and vary the probing dataset size \( n \). We then compute the generalization gap for each similarity metric. Figure 2 shows the simulation results, confirming that the generalization gap decays roughly as \( O(1/\sqrt{n}) \) for all metrics, matching our theoretical predictions.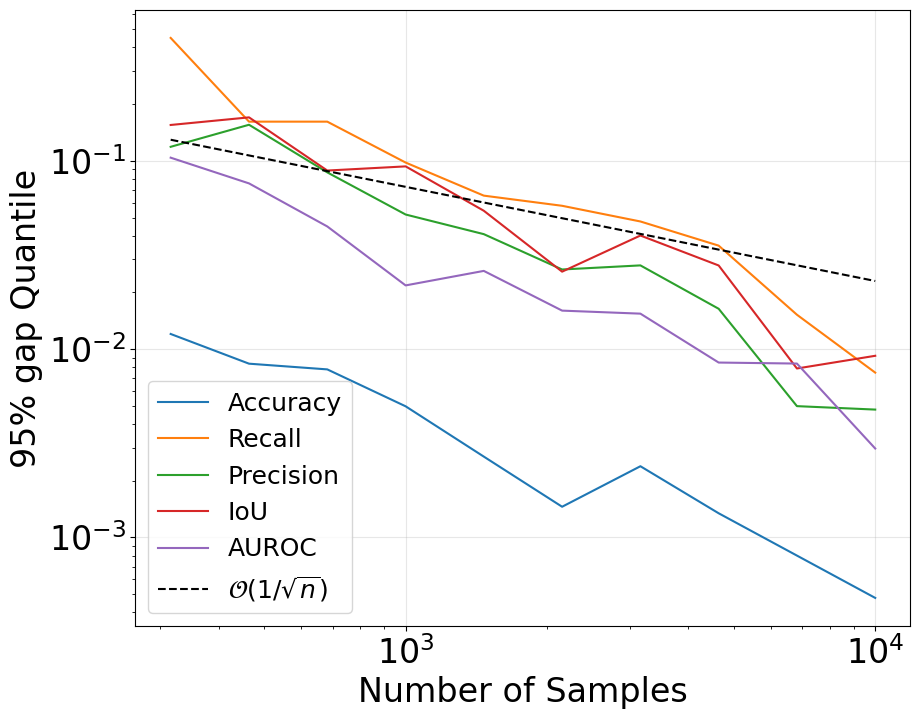
Figure 2: Simulated generalization gap decays roughly as \(O(1/\sqrt{n})\) for several metrics, matching the theory.
Contribution #2: Quantifying Stability via Bootstrap Explanation (BE)
(i) Overview
To assess whether the explanation would change under a different probing dataset, we propose a simple but general procedure inspired by the bootstrap ensemble method in machine learning:- Resample the probing dataset \(K\) times with replacement.
- Run the chosen neuron-identification method on each resampled dataset to get concepts \( \hat c_1, \dots, \hat c_K \).
- Aggregate counts to estimate a concept-level distribution.
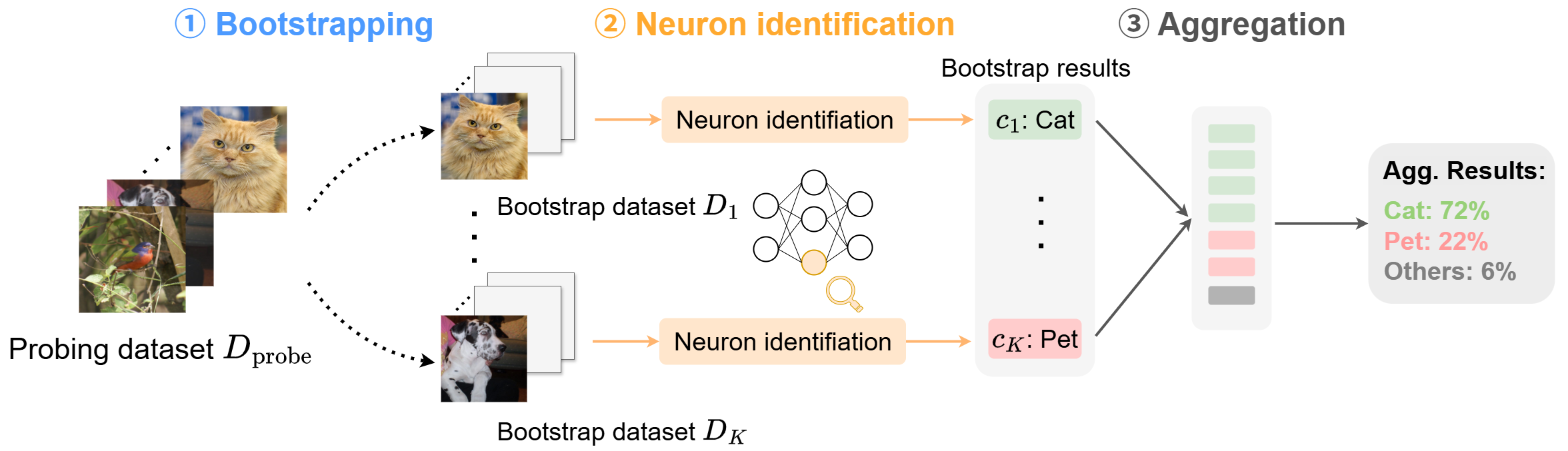
Figure 3: Bootstrap-based stability pipeline. Any existing neuron-identification method can be plugged in without modification.
Further, we develop the following algorithm to provide a set of potential concepts with statistical coverage guarantees:
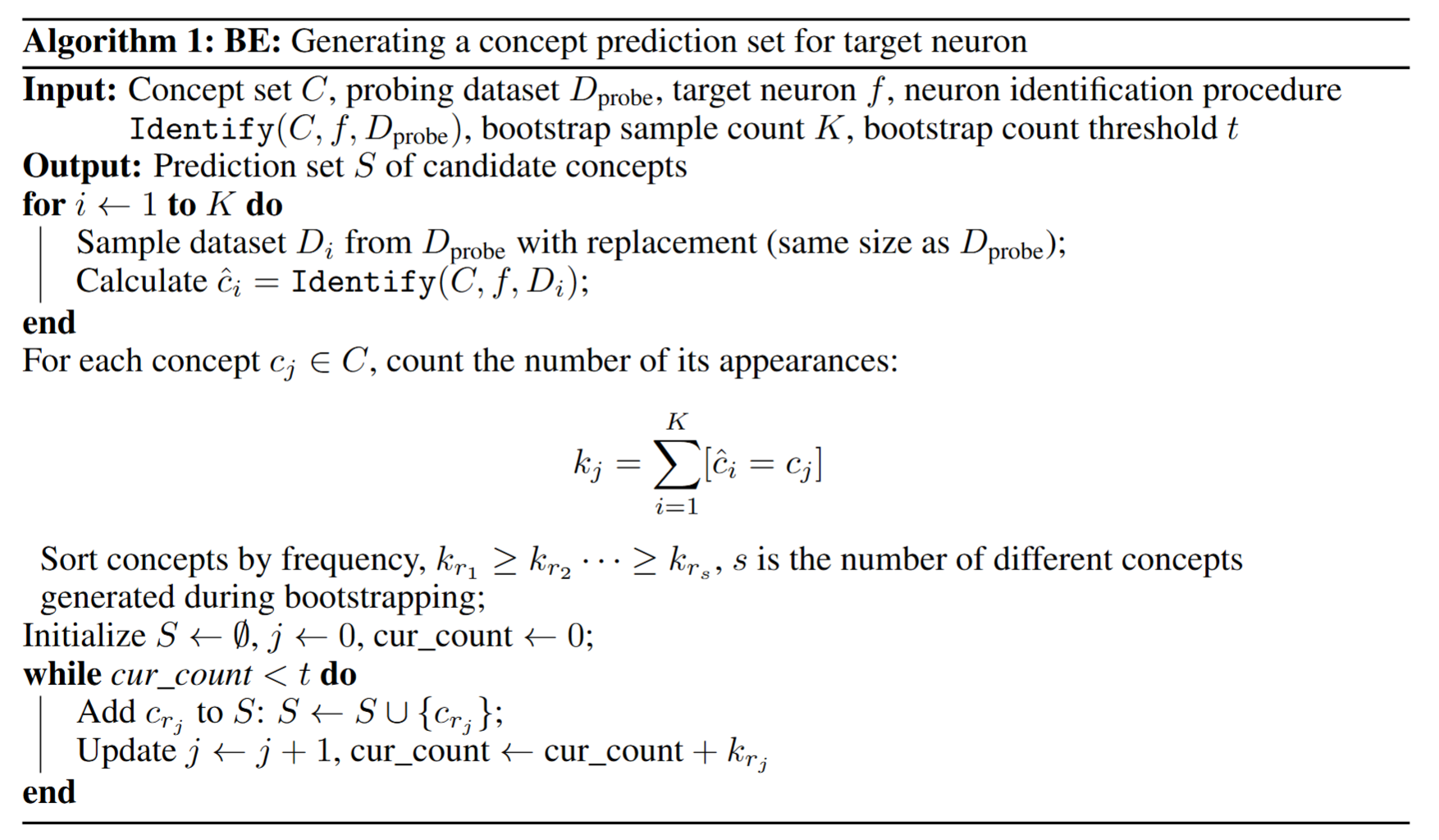
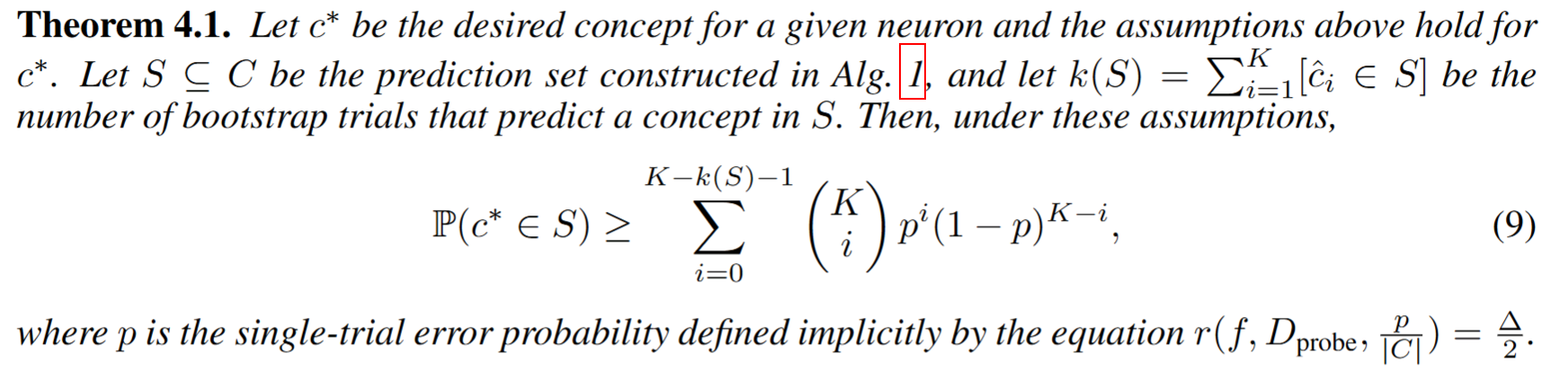


(ii) Experiments
We test our BE method on a ResNet-50 model using both NetDissect [1] and CLIP-Dissect [2].
Figure 4: Results of bootstrapping on two methods for ResNet-50 neurons.
References
[1] Bolei Zhou, David Bau, Aude Oliva, and Antonio Torralba. Network Dissection: Quantifying Interpretability of Deep Visual Representations. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017.
[2] Tuomas Oikarinen and Tsui-Wei (Lily) Weng. CLIP-Dissect: Automatic Description of Neuron Representations. ICLR, 2023.
[3] Tuomas Oikarinen, Ge Yan, and Tsui-Wei (Lily) Weng. Evaluating Neuron Explanations: A Unified Framework with Sanity Checks ICML, 2025.
Cite this work
Ge Yan, Tuomas Oikarinen, and Tsui-Wei (Lily) Weng. “Faithful and Stable Neuron Explanations for Trustworthy Mechanistic Interpretability.” NeurIPS Mechanistic Interpretability Workshop, 2025.
@article{
yan2025faithful,
title={Faithful and Stable Neuron Explanations for Trustworthy Mechanistic Interpretability},
author={Ge Yan and Tuomas Oikarinen and Tsui-Wei Weng},
booktitle={Mechanistic Interpretability Workshop at NeurIPS 2025},
year={2025},
url={https://openreview.net/forum?id=AxFsSMRu28}
}